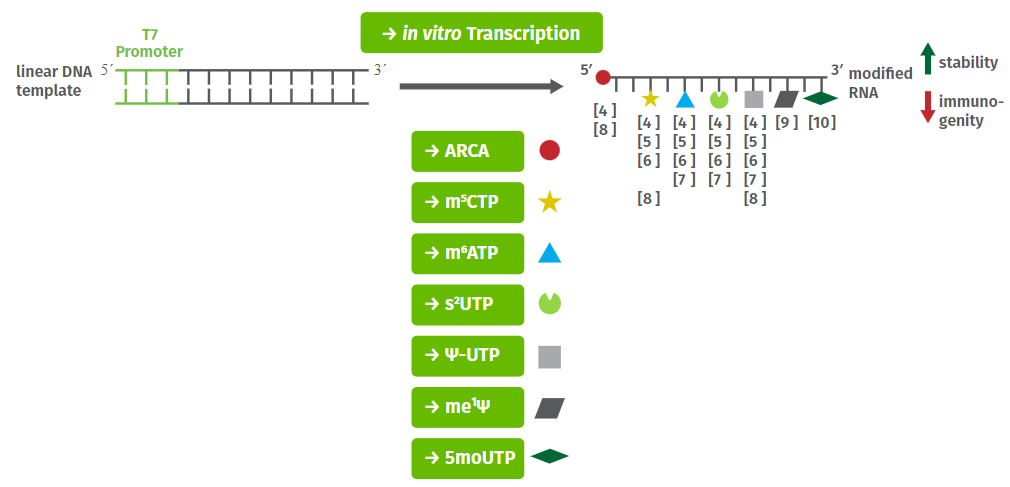
Nucleotides for SELEX/Aptamer Modification
Aptamers are short single-stranded DNA or RNA oligonucleotides (< 100 nt). They gained increasing importance in drug discovery due to their inherent ability to form defined three-dimensional structures that enables them to bind to various targets (e.g. proteins) with antibody-antigen-like affinity and specificity[1].
Aptamers with the highest binding affinity and specificity against a given target molecule are generated by an multi-step in vitro selection and enzymatic amplification process called “systematic evolution of ligands by exponential enrichment (SELEX) (for detailed information please refer to the references below[2,3]).
The main advantages of aptamers compared to antibodies are heat stability, the lack of immunogenicity and minimal interbatch variability. Their usage in diagnostics however, is often hampered by nuclease-mediated degradation.
Enzymatic incorporation of fluoro-, amino- or O-methyl- 2′-ribose-modified nucleotides (1) by generating an initially modified combinatorial library or (2) post-SELEX markedly improve the nuclease resistance of aptamers (Tab. 1) [3-12]. Furthermore, a number of modified nucleotides suitable for the selection of cross-linking capable aptamers (Photo-SELEX) are available as well (Tab. 1)[13-15].
Table 1: SELEX-compatible modified nucleotides.
n/a: not applicable
| Nucleotide | Modification | DNA aptamer selection (DNA SELEX) |
RNA aptamer selection (RNA SELEX) |
|
| Ribose Moiety | 2’F-dUTP | Substitution of 2′-OH by fluor (F) | [5] | [6,7] |
| 2’F-dCTP | [5] | [6,7] | ||
| 2’F-dATP | [5] | [6] | ||
| 2’F-dGTP | [5] | |||
| 2’OMe-UTP | Modification of 2′-OH by a methyl group (CH3) | [8,9,10,11] | ||
| 2’OMe-CTP | [8,9,11] | |||
| 2’OMe-ATP | [8] | |||
| 2’OMe-GTP | [8] | |||
| 2’NH2-dUTP | Substitution of 2′-OH by an amino group (NH2) | [6,12] | ||
| 2’NH2-dCTP | [12] | |||
| 2’NH2-dATP | ||||
| 2’NH2-dGTP | ||||
| LNA-ATP | LNA (Locked Nucleic Acid) with methylene bridge beetween 2′-O and 4′-C | [16] | ||
| LNA-GTP | ||||
| LNA-CTP | ||||
| LNA-UTP | ||||
| Base Moiety | 5Br-dUTP | Modification of C-5 by Bromine (Br) | [13] | n/a |
| 5I-UTP | Modification of C-5 by Iodine (I) | n/a | [14] | |
| 4-Thio-UTP | Substitution of 4-O by Sulfur (S) | n/a | [15] |
| Name | Cat. No. | Size |
| 2′-Fluoro-dUTP | NU-1215S | 50 μl (100 mM) |
| 2′-Fluoro-dUTP | NU-1215L | 5 x 50 μl (100 mM) |
| 2′-Fluoro-dCTP | NU-1214S | 50 μl (100 mM) |
| 2′-Fluoro-dCTP | NU-1214L | 5 x 50 μl (100 mM) |
| 2′-Fluoro-dATP | NU-151S | 50 μl (100 mM) |
| 2′-Fluoro-dATP | NU-151L | 5 x 50 μl (100 mM) |
| 2′-Fluoro-dGTP | NU-1216S | 50 μl (100 mM) |
| 2′-Fluoro-dGTP | NU-1216L | 5 x 50 μl (100 mM) |
| 2′-Fluoro-dNTP Bundle | NU-10405-20 | 4 x 50 μl (4 x 5 μmol) |
| 2’OMe-UTP | NU-1212S | 50 μl (100 mM) |
| 2’OMe-UTP | NU-1212L | 5 x 50 μl (100 mM) |
| 2’OMe-CTP | NU-1211S | 50 μl (100 mM) |
| 2’OMe-CTP | NU-1211L | 5 x 50 μl (100 mM) |
| 2’OMe-ATP | NU-1184S | 50 μl (100 mM) |
| 2’OMe-ATP | NU-1184L | 5 x 50 μl (100 mM) |
| 2’OMe-GTP | NU-1127S | 50 μl (100 mM) |
| 2’OMe-GTP | NU-1127L | 5 x 50 μl (100 mM) |
| 2’NH2-dUTP | NU-242S | 50 μl (100 mM) |
| 2’NH2-dUTP | NU-242L | 5 x 50 μl (100 mM) |
| 2’NH2-dCTP | NU-243S | 50 μl (100 mM) |
| 2’NH2-dCTP | NU-243L | 5 x 50 μl (100 mM) |
| 2’NH2-dATP | NU-244S | 50 μl (100 mM) |
| 2’NH2-dATP | NU-244L | 5 x 50 μl (100 mM) |
| 2’NH2-dGTP | NU-245S | 50 μl (100 mM) |
| 2’NH2-dGTP | NU-245L | 5 x 50 μl (100 mM) |
| LNA-ATP | NU-982 | 10 μl (100 mM) |
| LNA-GTP | NU-983 | 10 μl (100 mM) |
| LNA-CTP | NU-984 | 10 μl (100 mM) |
| LNA-UTP | NU-985 | 10 μl (100 mM) |
| 5-Bromo-dUTP | NU-122S | 50 μl (10 mM) |
| 5-Bromo-dUTP | NU-122L | 5 x 50 μl (10 mM) |
| 5-Iodo-UTP | NU-119S | 10 μl (100 mM) |
| 5-Iodo-UTP | NU-119L | 5 x 10 μl (100 mM) |
| 4-Thio-UTP | NU-1156S | 10 μl (100 mM) |
| 4-Thio-UTP | NU-1156L | 5 x 10 μl (100 mM) |