CRISPR/Cas9
Genome Engineering Tools from Novabiotein
Cas9 (CRISPR associated protein 9) is an RNA-guided DNA endonuclease enzyme associated with the CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats) adaptive immunity system in Streptococcus pyogenes. S. Pyogenes utilizes Cas9 to remember and later probe and cleave foreign DNA, such as invading bacteriophage DNA or plasmid DNA. When the DNA substrate is complementary to the guide RNA, Cas9 cleaves the invading DNA. This ability to make site-directed double strand breaks in DNA has led to the use of Cas9 protein as a genome engineering tool. As an RNA-guided DNA endonuclease, Cas9 can be directed to specific target cleavage sites by altering the guide RNA sequence. The nuclease-deactivated form of dCas9 further provides a versatile tool for biomedical research and therapeutics.
Stratech Scientific offers a comprehensive range of CRISPR/Cas9 related enzymes, proteins, antibodies and kits for genome engineering from Novateinbio.
Transfection-ready Cas9 proteins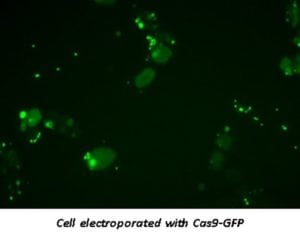
Compared to transfection with Cas9-expressing plasmid, delivery of purified Cas9 protein directly into cells by electroporation is relatively straightforward and may reduce the risk of issues associated with plasmid-based transfection, such as random integration of plasmid into the host genome. The presence of a nuclear localization signal (NLS) localizes the Cas9 RNP complex to the nucleus immediately upon entering the cell with the need for transcription or translation compared with mRNA or plasmid systems. Additionally, the Cas9 RNP complex is rapidly cleared from the cells minimizing the chance of off-target cleavage when compared to other systems. There is no need for transcription or translation processes.
NLS-Cas9-NLS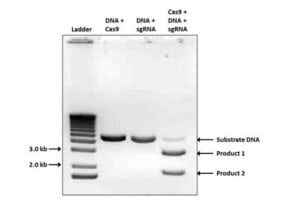
Cas9 NLS (PR-137211), is a recombinant dCas9 from S. pyogenes which contains Simian virus 40(SV40) T antigen nuclear localization sequence (NLS) on the N- and C- termini of the protein and is used to create double blunt end for recombination. The protein is transfection-ready for gene editing, also used for sgRNA or crRNA screening.
NLS-dCas9-NLS
NLS-dCas9-NLS (PR-137213B) is an engineered, nuclease- deactivated Cas9 (dCas9) protein, has been used to regulate endogenous gene expression and labelling of genomic loci in living and fixed cells. The protein has been mutated CRISPR-associated endonuclease Cas9 (amino acids 1 to 1368) with D10A & H840A (ACCESSION: AKS40378 for Cas9). To facilitate nuclear entry, two nuclear localization signal sequence (NLS) are fused to both N- and C-terminal of dCas9 protein.
NLS-Cas9-EGFP
for subcellular localization of the Cas9-gRNA/target DNA complex.
NLS-Cas9-EGFP (PR-137211E) is a fusion protein containing a nuclear localization sequence (NLS) on its N terminal and EGFP on the C terminal. The EGFP can be taken as a reporter for tracking or sorting transfected cells, to enrich for cell populations containing the desired genome edits by fluorescence activated cell sorting (FACS). It significantly reduces the labour and cost associated with single cell cloning and genotyping in genome editing applications.
NLS-Cas9(D10A)-2NLS nickase
NLS-Cas9(D10A)-2NLS nickase (PR-137212B) is a transfection-ready, recombinant Cas9 for gene editing, single nick creation on double strand DNA. NLS-Cas9-D10A Nickase is a mutation form of Cas9 Nuclease which makes one active domain deactivated, thus it can only cut one single strand DNA that is complementary to the guide- RNA, producing one single strand cut. Combined with two different gRNA, NLS-Cas9-D10A Nickase produces two cut sites respectively and causes a double strand break. Compared with the wild type Cas9 nuclease, the two- gRNA guided cleavage can significantly reduce the off-target effects.
Screening Tools for CRISPR/Cas9
sgRNA Screening Kit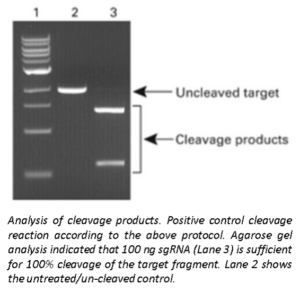
The sgRNA Screening Kit (NB-GEN10642) is designed to quickly assess in vitro the potential efficiency of a sgRNA sequence in vivo. The design of single guide RNA (sgRNA) is dependent on the target region close to the PAM site. Because the specificity and activity of each sgRNA is unpredictable, it is often recommended to design multiple sgRNAs to target a gene of interest. This kit provides a simple, reliable, and rapid in vitro method to assess single guide RNA (sgRNA) efficiency before cell transformation, to identify the most effective CRISPR sgRNA from a pool of sgRNAs, in as little as 2 hours.
Anti-CRISPR/CAS9 Monoclonal Antibody
This IgG1kappa mouse monoclonal antibody (SH-A16960) is specific for Cas9 ( wild type, nickase and dCas9) from all species using Western Blot, Immunofluorescence, and ELISA.
Anti-CRISPR/CAS9 Polyclonal Antibody
This rabbit polyclonal IgG (SH-A16959) recognizes both Cas9 and dCas9 (nuclease-deactivated Cas9) based on the antigen design. It is suitable for us in ELISA,Western Blot,Immunofluorescnece, ChIP, Immunoprecipitation assays.
CRISPR/Cas9 Protein ELISA Kit (Colorimetric)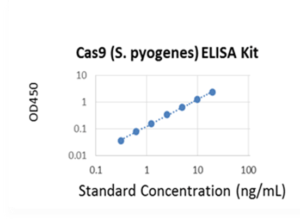
The CRISPR/Cas9 Protein ELISA Kit (Colorimetric) (NB-E1372PR) is designed for measuring Cas9 level in whole cell extracts or purified Cas9 proteins from various cells and tissues. In an assay with this kit, the Cas9 proteins in samples are tightly and stably spotted on the strip wells. The bound Cas9 proteins can then be recognized with detection antibody followed by a colour development reagent. The ratio of Cas9 is proportional to the intensity of absorbance. The absolute amount of Cas9 can be quantitated by comparing to the Cas9 control.
References
1. Deactivated CRISPR Associated Protein 9 for Minor-Allele Enrichment in Cell-Free DNA Amin Aalipour et al. Clinical Chemistry 2018 Vol. 64, Issue 2 p307-p916
2. Purified Cas9 Fusion Proteins for Advanced Genome Manipulation Jovan Mircetic et al. Small Methods 2017 1, 1600052
3. Disruptive non-disruptive applications of CRISPR/Cas9 Jonathan LSchmid-Burgk Current Opinion in Biotechnology December 2017, Volume 48 Pages 203-209
4. Efficient sequence-specific isolation of DNA fragments and chromatin by in vitro enChIP technology using recombinant CRISPR ribonucleoproteins Toshitsugu Fujita et al. Genes to Cells Volume 21, Issue 4 April 2016 Pages 370–377
5. High-throughput biochemical profiling reveals sequence determinants of dCas9 off-target binding and unbinding Evan A Boyle et al. PNAS 2017 May, 114 (21) 5461-5466
6. CASFISH: CRISPR/Cas9-mediated in situ labeling of genomic loci in fixed cells Wulan Deng et al. PNAS 2015 vol. 112 | no. 38 p11870-11875
Streptococcus pyogenes Cas9 variants used for genome editing from Novabiotein
| Product name | Catalog number | CRISPR-Cas9 variants | Amino acid features | NLS position | Applications | PAM sequence |
| Cas9 nuclease | PR-137210 | Streptococcus pyogenesCas9 wild type | wild type | N-terminal | Create double blunt end for recombination | NGG |
| NLS-Cas9-NLS | PR-137211 | Streptococcus pyogenesCas9 wild type | wild type | N-terminal and C-terminal | Create double blunt end for recombination | NGG |
| NLS-Cas9-EGFP | PR-137211-E | Streptococcus pyogenesCas9 with C-terminal EGFP | wild type | N-terminal and C-terminal | Create double blunt end for recombination | NGG |
| Cas9(D10A) nickase | PR-137212 | Streptococcus pyogenesCas9 nickase | D10A | N-terminal | Create nick on one strand | NGG |
| NLS-Cas9(D10A)-2NLS nickase | PR-137212B | Streptococcus pyogenesCas9 nickase | D10A | N-terminal and 2xC-terminal | Create nick on one strand | NGG |
| dCas9 (D10A & H840A) | PR-137213 | no nuclease activity while keeping target binding activity | D10A/H840A | C-terminal | Detection and purification targeted DNA | NGG |
| NLS-dCas9-NLS | PR-137213B | no nuclease activity while keeping target binding activity | D10A/H840A | N-terminal and C-terminal | Detection and purification targeted DNA | NGG |
| Biotinylated NLS-dCas9-NLS | PR-137213-bio | no nuclease activity while keeping target binding activity, biotinylated | D10A/H840A | N-terminal and C-terminal | Specific DNA fragment pull-down in CHIP assay. FISH and short sequence dtection | NGG |
| NLS-dCas9-EGFP-NLS | PR-137213G | no nuclease activity while keeping target binding activity | D10A/H840A | N-terminal and C-terminal | FISH, genomic visualization | NGG |
| eSpCas9(1.1)-3NLS | Cas9-HiFi1 | Cas9 with increase On-target specificity | K848A/K1003A/R1060A | N-terminal and 2xC-terminal | gene editing with better on-target, less off-target rate. | NGG |
| SpCas9-HF1-3NLS | Cas9-HiFi2 | Cas9 with increase On-target specificity | N497A/R661A/Q695A/Q926A | N-terminal and 2xC-terminal | gene editing with better on-target, less off-target rate. | NGG |