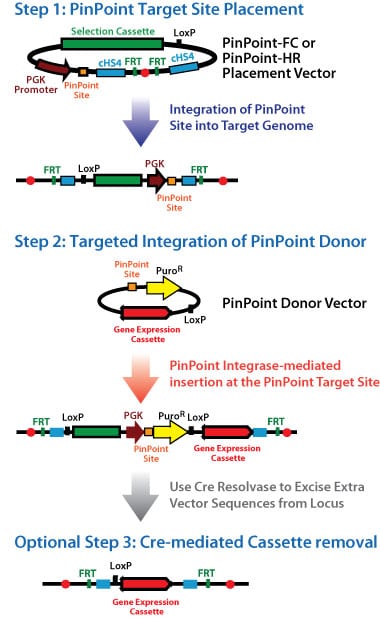
How the PinPoint system works
The PinPoint system is a two-step approach for engineering of target cells with an optional third step for selection cassette removal by Cre resolvase. The first step involves insertion of a plasmid bearing the PinPoint placement site via transfection into the target cell genome. This can be done using two distinct approaches: i) the PinPoint-FC system that involves the well-characterized phiC31 integrase system or ii) the PinPoint-HR system that uses either TALE-Nuclease or Cas9 genome engineering tools to induce a double-stranded DNA break in the genome and insertion of the PinPoint placement site by homology-directed recombination (HDR) in a site-specific manner.
The second part of the PinPoint system relies on the introduction of a donor vector containing your desired gene cassette insert, which is integrated into the placed PinPoint site using a hyperspecific and efficient PinPoint integrase. The PinPoint integrase catalyzes the attB x attP reaction between the placed site (attP) and the attB site in the donor vector to insert the donor vector at the placed site each and every targeting event.
The third optional step invloves the removal of the entire backbone (excluding the insert and its promoter) using the well-characterized Cre/LoxP reaction leaving only the promoter/insert combination (and a single LoxP site) in the genome.
Target site placement with the PinPoint-FC system
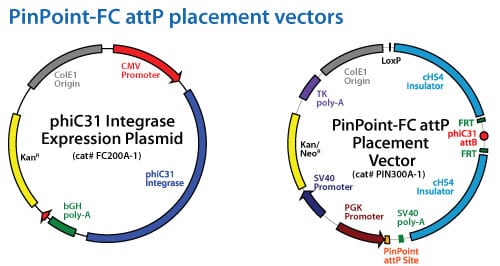
Platform cell lines containing the PinPoint attP site can be made through two different approaches. The PinPoint-FC system utilizes the phiC31 integrase (Cat# FC200A-1) along with the PinPoint-FC attP Placement Vector (Cat# PIN300A-1). This will integrate the cassette at pseudo attP sites within the targeted genome with typically single copy insertions.
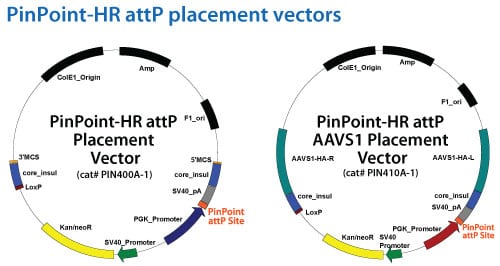
Target site placement with the PinPoint-HR system
Alternatively, the PinPoint-HR system uses a site-directed placement strategy. Customized placement can be achieved using the PIN400A-1 HR vector. First, clone the homology arms for the desired integration site into the 5′ and 3′ MCS features. Then use this vector in combination with matched TALENs or Cas9 nuclease vectors.
SBI has created a safe-harbor PinPoint placement vector (Catalogue no. PIN410A-1) to insert the attP site at the human AAVS1 genomic locus for routine integration of gene expression cassettes at this site.
PinPoint site targeting
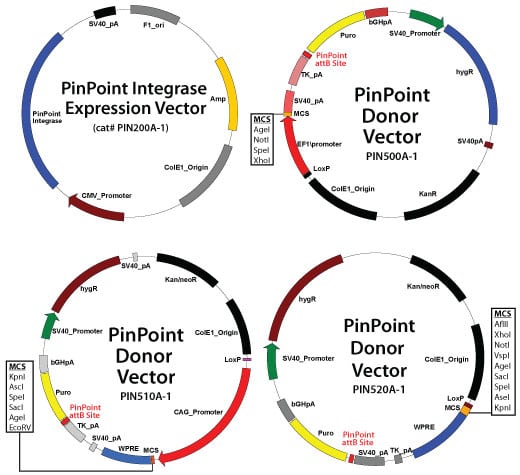
Once the PinPoint attP site has been successfully placed into the desired genomic site, this site can then be efficiently and routinely targeted for site-specific integration of expression cassettes at that locus. The expression cassette can be driven by EF1a (Cat# PIN500A-1), CAGs (Cat# PIN510A-1), or a custom-designed promoter of your choice (Cat# PIN520A-1). Once the gene or reporter of interesthas been cloned into the PinPoint donor vector, a simple co-transfection of the placed attP site cell line with the PinPoint integrase vector (Cat# PIN200A-1) will result in a specific recombination reaction between the PinPoint attP site and the attB site in the donor vector construct.
PinPoint integrase and donor vectors
.
PinPoint site placement and targeting
Pinpoint-FC attP placement
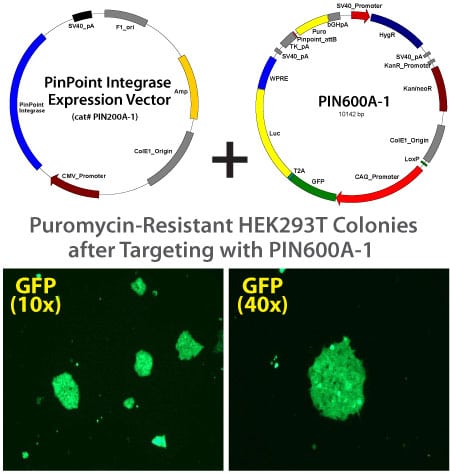
The PinPoint attP site was efficiently placed into HEK293 cells or HeLa cells. Approximately 8×10^5 cells were transfected with 40 ng PinPoint-FC placement vector (Cat# PIN300A-1) and 1,960 ng phiC31 integrase expression vector (Cat# PIN200A-1) at a 1:50 ratio, respectively. Positive cells were selected with 400 ug/ml G418 for 14 days. Six separate lines of HEK293 and HeLa cells were picked and expanded. After 11 days, the cells were fixed and stained with a solution of 50% methanol plus 1% methylene blue. The plates were washed twice with PBS and allowed to air dry. The number of visible colonies were imaged and colony number counts assessed.
Neomycin resistant HEK293 cells with placed PinPoint site
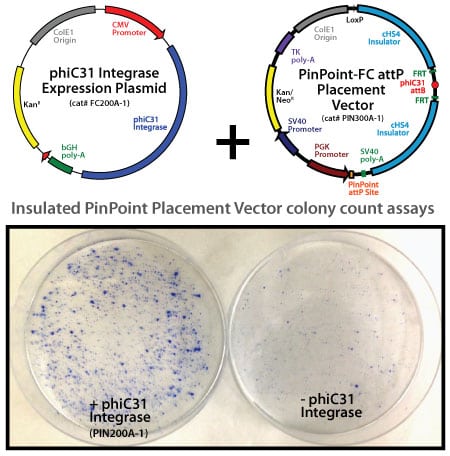

These lines were then transfected with a control PinPoint Gene Donor vector (PIN600A-1) for precise integration. Unlike the R4 integrase used in a similar system, the PinPoint integrase does not recognize pseudo-sites and will only integrate at its placed recognition sequence. This feature results in the increased efficiency of correctly retargeted cell lines compared to the R4 integrase system. The entire therapeutic gene expression cassette is active and fully insulated with cHS4 DNA elements.
Neomycin resistant HEK293 cells with placed PinPoint site
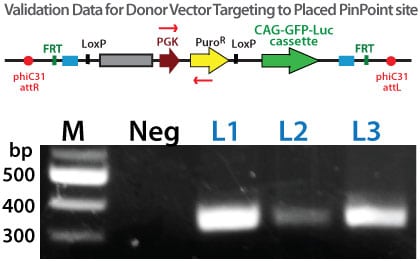
To verify that the PinPoint donor construct targeted to the correct site, junction PCR can be employed to amplify across the PinPoint integrase recombination site (attL) junction. Primers are designed within the placed puromycin marker cassette and within the PGK promoter are used in nested PCR reactions to verify the correct PinPoint integration structure. Three separate HEK293 colonies that were GFP positive were expanded for 3 days with subsequent genomic DNA isolation. Approximately 50 ng of genomic DNA was used in the nested PCR reactions and the products separated on 2% agarose gels and stained with ethidium bromide before imaging. The results clearly indiavcte the predicted, PinPoint integrase-mediated insertion of the PIN600a-1 donor vector at the placed attP site with high precision.
Online Resources
- PinPoint Integrase System User Manual (PDF)
Technical References
- Karow M, Chavez CL, Farruggio AP, Geisinger JM, Keravala A, Jung WE, Lan F, Wu JC, Chen-Tsai Y, Calos MP. Site-specific recombinase strategy to create induced pluripotent stem cells efficiently with plasmid DNA. Stem Cells 29(11):1696-704.
- Chalberg TW, Portlock JL, Olivares EC, Thyagarajan B, Kirby PJ, Hillman RT, Hoelters J, Calos MP. Integration specificity of phage phiC31 integrase in the human genome. J Mol Biol. 2006 Mar 17;357(1):28-48.
- Lan F, Liu J, Narsinh KH, Hu S, Han L, Lee AS, Karow M, Nguyen PK, Nag D, Calos MP, Robbins RC, Wu JC. Safe genetic modification of cardiac stem cells using a site-specific integration technique. Circulation. 2012 Sep 11;126(11 Suppl 1):S20-8.
- Hillman RT, Calos MP. Site-specific integration with bacteriophage phiC31 integrase. Cold Spring Harb Protoc. 2012 May 1;2012(5).
- Chavez CL, Keravala A, Chu JN, Farruggio AP, Cuellar VE, Voorberg J, Calos MP. Long-term expression of human coagulation factor VIII in a tolerant mouse model using the phiC31 integrase system. Hum Gene Ther. 2012 Apr;23(4):390-8.
- Woodard LE, Keravala A, Jung WE, Wapinski OL, Yang Q, Felsher DW, Calos MP. Impact of hydrodynamic injection and phiC31 integrase on tumor latency in a mouse model of MYC-induced hepatocellular carcinoma. PLoS One. 2010 Jun 29;5(6):e11367.
- Carr PA, Church GM. Genome engineering. Nat Biotechnol. 2009 Dec;27(12):1151-62.
- Bhaya D, Davison M, Barrangou R. CRISPR-Cas systems in bacteria and archaea: versatile small RNAs for adaptive defense and regulation. Annu Rev Genet. 2011;45:273-97.
- Terns MP, Terns RM. CRISPR-based adaptive immune systems. Curr Opin Microbiol. 2011 Jun;14(3):321-7.
- Makarova KS, Haft DH, Barrangou R, Brouns SJ, Charpentier E, Horvath P, Moineau S, Mojica FJ, Wolf YI, Yakunin AF, van der Oost J, Koonin E. Evolution and classification of the CRISPR-Cas systems. Nat Rev Microbiol. 2011 Jun;9(6):467-77.
- Wiedenheft B, Sternberg SH, Doudna JA. RNA-guided genetic silencing systems in bacteria and archaea. Nature. 2012 Feb 15;482(7385):331-8.
- Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science. 2012 Aug 17;337(6096):816-21.
- Mali P, Yang L, Esvelt KM, Aach J, Guell M, DiCarlo JE, Norville JE, Church GM. RNA-guided human genome engineering via Cas9. Science. 2013 Feb 15;339(6121):823-6.
- Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, Hsu PD, Wu X, Jiang W, Marraffini LA, Zhang F. Multiplex genome engineering using CRISPR/Cas systems. Science. 2013 Feb 15;339(6121):819-23.
- Jinek M, East A, Cheng A, Lin S, Ma E, Doudna J. RNA-programmed genome editing in human cells. Elife. 2013;2:e00471. doi: 10.7554/eLife.00471.
PinPoint Targeted Integration System
| Product | Size | Catalogue No. |
| PinPoint Integrase Expression Plasmid | 10 ug | PIN200A-1 |
| PinPoint-FC attP Placement Vector | 10 ug | PIN300A-1 |
| PinPoint-FC System for Platform Cell Line Generation & Retargeting (includes PIN300A-1, FC200PA-1, PIN200A-1, PIN510A-1, & PIN600A-1) | 1 Kit | PIN300A-KIT |
| PinPoint-FC 293T Platform Cell Line for Targeted Gene Insertion (with PinPoint site already placed) | >2×10^5 cells | PIN320A-1 |
| PinPoint-FC 293T Platform Kit for Targeted Gene Insertion (includes PIN320A-1, PIN200A-1, PIN510A-1 & PIN600A-1) | 1 Kit | PIN320A-KIT |
| PinPoint-FC Murine iPSC Platform Cell Line for Targeted Gene Insertion (with PinPoint site already placed) | >2×10^5 cells | PIN340iPS-1 |
| PinPoint-FC Murine iPSC Platform Kit for Targeted Gene Insertion (includes PIN340iPS-1, PIN200A-1, PIN510A-1 & PIN600A-1) | 1 Kit | PIN340iPS-KIT |
| PinPoint-HR attP Placement Vector Empty MCS for cloning HR Arms | 10 ug | PIN400A-1 |
| PinPoint-HR System for Platform Cell Line Generation & Retargeting (includes PIN400A-1, PIN200A-1, PIN510A-1, & PIN600A-1) | 1 Kit | PIN400A-KIT |
| PinPoint-HR attP Placement Vector for targeting the Human AAVS1 Safe Harbor | 10 ug | PIN410A-1 |
| PinPoint-HR System for Platform Cell Line Generation & Retargeting of AAVS1 Safe Harbor Locus (includes PIN410A-1, GE601A-1, PIN200A-1, PIN510A-1, & PIN600A-1) | 1 Kit | PIN410A-KIT |
| PinPoint-HR System for Platform Cell Line Generation & Retargeting of AAVS1 Safe Harbor Locus (includes PIN410A-1, CAS601A-1, PIN200A-1, PIN510A-1, & PIN600A-1) | 1 Kit | PIN412A-KIT |
| EF1alpha PinPoint Donor Vector (pPP-EF1a-MCS-pA-attB-Puro-pA) | 10 ug | PIN500A-1 |
| CAG PinPoint Donor Vector (pPP-CAG-MCS-WPRE-pA-attB-Puro-pA) | 10 ug | PIN510A-1 |
| Empty PinPoint Donor Vector (pPP-MCS-WPRE-pA-attB-Puro-pA) | 10 ug | PIN520A-1 |
| PinPoint Donor Positive Control Vector: CAG->GFP+Luciferase (pPP-CAG-GFP-T2A-Luc-WPRE-pA-attB-Puro-pA) | 10 ug | PIN600A-1 |
| PinPoint Donor Positive Control Vector: CM->Luciferase (pPP-CMV-Luc-pA-attB-Puro-pA) | 10 ug | PIN610A-1 |
| Cre Recombinase Expression Plasmid (pCMV-CRE) | 10 ug | CRE100A-1 |
| PhiC31 Integrase Expression Plasmid | 10ug | FC200PA-1 |
| AAVS1 Safe Harbor TALE Nuclease pair [pZT-AAVS1-L1 and pZT-AAVS1-R1] | 10 ug | GE601A-1 |
| Cas9 SmartNuclease AAVS1-gRNA Targeting Vector [EF1a promoter] | 10 ug | CAS601A-1 |