RNA sequencing (RNA-seq) provides a molecular snapshot of the cellular transcriptome, offering a treasure trove of information regarding gene regulation and expression. One of the most common—and arguably the most biologically relevant—applications of RNA sequencing is the ability to assess differential gene expression (DGE), which provides invaluable data on genome-wide differences in gene expression between biological samples. As changes in gene expression often involve multiple comparisons—such as between wild types and mutants, life history stages, experimental time points, or environmental conditions—and normally include controls and replicates, DGE analyses often involve numerous RNA samples. However, until recently, traditional RNA-seq strategies have been too laborious, expensive, and analytically challenging to replace reverse transcription-quantitative real-time PCR (qRT-PCR) as the preferred method of quantifying changes in gene expression, thereby limiting the number of RNA samples that can be processed (Alpern et al. 2019).
The advent of Bulk RNA Barcoding and sequencing (BRB-seq)—an innovative, efficient, and cost-effective RNA sequencing technology—has revolutionized RNA-seq workflows (Alpern et al. 2019). BRB-seq, which applies sample barcoding and early multiplexing principles to bulk-RNA profiling, enables large numbers of samples to be pooled and sequenced simultaneously.
Figure 1. Each well of the BRB-seq oligo(dT) primer plate contains a distinct barcode adaptor to uniquely label the RNA sample added to each well. After reverse transcription, barcoded samples are pooled for the remainder of the cDNA library preparation workflow and library sequencing.
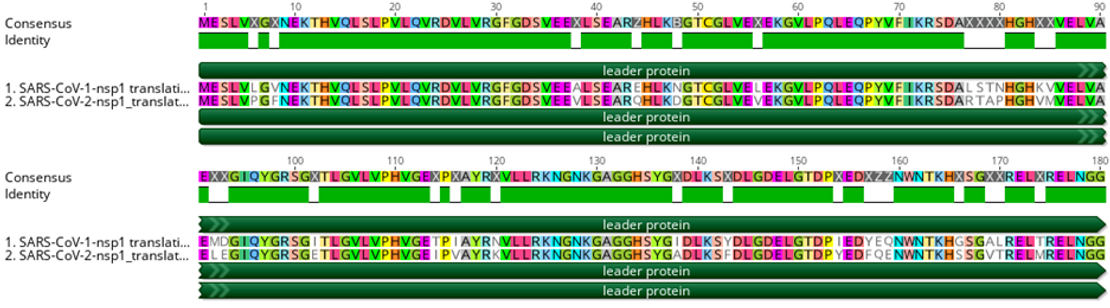
As incubation temperature has been previously shown to affect replication kinetics in coronaviruses and other respiratory viruses (Hoorn and Tyrrell 1966; Corman et al. 2016), a recent study by V’kovski et al. (2021) used BRB-seq to assess DGE between coronaviruses SARS-CoV-1 and SARS-CoV-2 at two biologically relevant temperatures (33 and 37°C) to mimic the upper and lower respiratory tracts, respectively. Despite their phylogenetic similarities, most notably in functionally conserved regions (Figure 2), SARS-CoV-1 and SARS-CoV-2 are known to exhibit considerable differences in human-to-human transmission dynamics, pathogenicity, and clinical courses of infection (Cevik, Kuppalli, et al. 2020; Cevik, Tate, et al. 2020), indicating possible differences in virus-host dynamics (V’kovski et al. 2021).
Figure 2. An amino acid alignment of SARS-CoV-1 and SARS-CoV-2 non-structural protein 1 (i.e., leader protein). Green rectangles below the consensus sequence represent amino acids that are identical between the two sequences.
To explore potential differences in virus-host dynamics and elucidate the effect of incubation temperature on the viral replication kinetics and host immune response of SARS-CoV-1 and SARS-CoV-2, the authors infected human airway epithelial cell (hAEC) cultures with 30,000 plaque-forming units of either SARS-CoV-1 or SARS-CoV-2 and incubated the infected cultures at either 33 or 37°C to mimic respiratory tract conditions. In addition, some cultures were pretreated with exogenous type I and III interferon (IFN) to assess the innate immune response to viral infection prior to incubation at the experimental temperatures. BRB-seq was used to assess DGE in response to the experimental conditions every 24 h for up to 96 h post infection for both viruses using released virus progeny.
The authors found that while SARS-CoV-1 and SARS-CoV-2 replicated to similar titers over the course of infection at 37°C (i.e., lower airway), SARS-CoV-2 infection resulted in 10-fold higher titers at 33°C (i.e., upper airway) between 72 and 96 h post infection. Furthermore, viral replication was attenuated in both viruses after incubation with interferons at both experimental temperatures. Given the substantial effect of incubation temperature on replication kinetics in SARS-CoV-2, the authors sought to investigate the effects of the two experimental temperatures on the host transcriptional response to SARS-CoV-1 and SARS-CoV-2 at 24, 48, 72, and 96 h post infection using BRB-seq. Samples were sequenced at a depth of 12 million reads per sample to ensure that genes with lower levels of expression were captured in the analysis. BRB-seq allowed for the downstream analysis of three distinct approaches to DGE: i) a comparison of DGE between SARS-CoV-1 and SARS-CoV-2 in response to temperature, ii) pairwise comparisons between each virus and control cultures at each timepoint, and iii) an analysis of changes in gene expression over time.
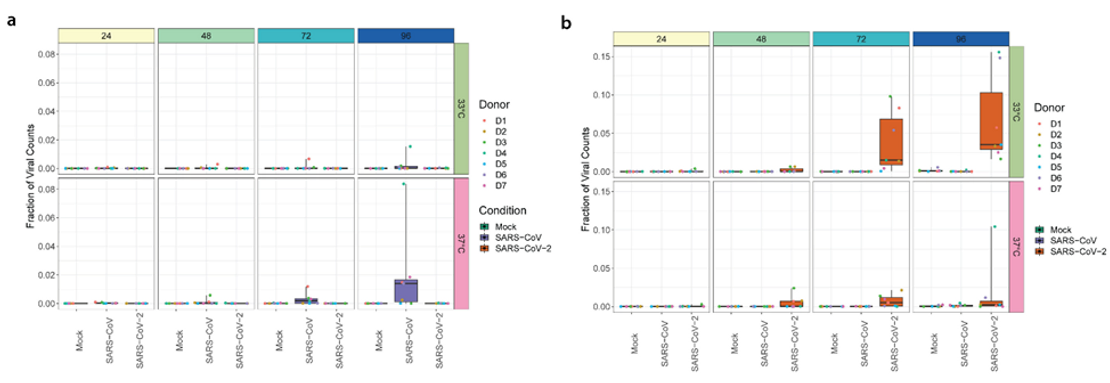
Figure 3. Boxplots illustrating the distribution of sequencing reads attributed to a) SARS-CoV-1 and b) SARS-CoV-2 over time for each temperature condition over time. Figure adapted from V’kovski et al. (2021).
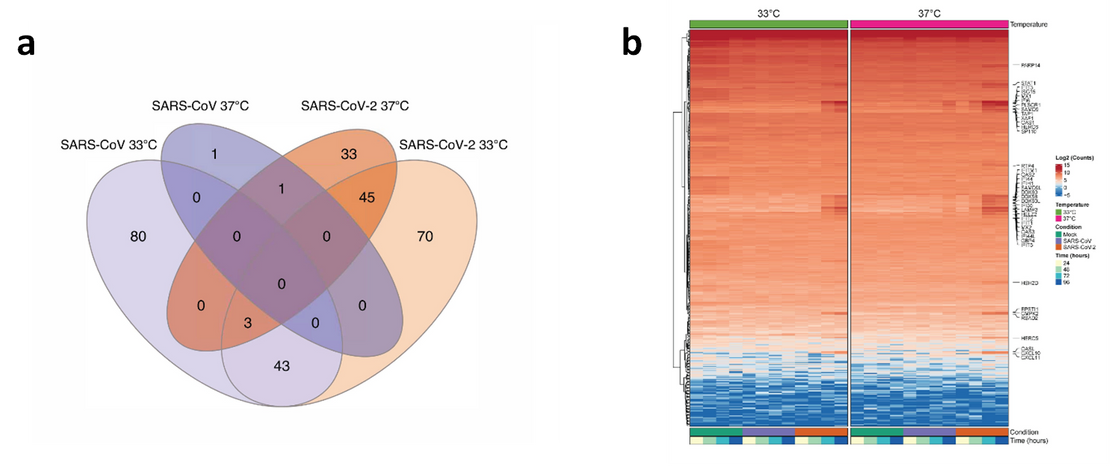
The results of BRB-seq analyses show that the quantities of SARS-CoV-1 and SARS-CoV-2 specific sequence reads were consistent with the results of viral replication kinetics (Figure 3). For the second approach, BRB-seq enabled the identification of 126 DEGs for SARS-CoV-1 at 33°C and 2 DEGs at 37°C, while 161 DEGs were identified for SARS-CoV-2 at 33°C and 82 DEGs at 37°C, for a total of 276 uniquely expressed genes. Lastly, a comparison of the DEGs for each virus at each temperature revealed gene clusters that were both shared and distinctly associated with each virus (Figure 4a). A hierarchical cluster analysis of DEGs at the two experimental temperatures for each virus revealed several distinct gene clusters that were specific to either SARS-CoV-1 or SARS-CoV-2 in comparison to control cultures and others that were consistent between experimental conditions and timepoints (Figure 4b).