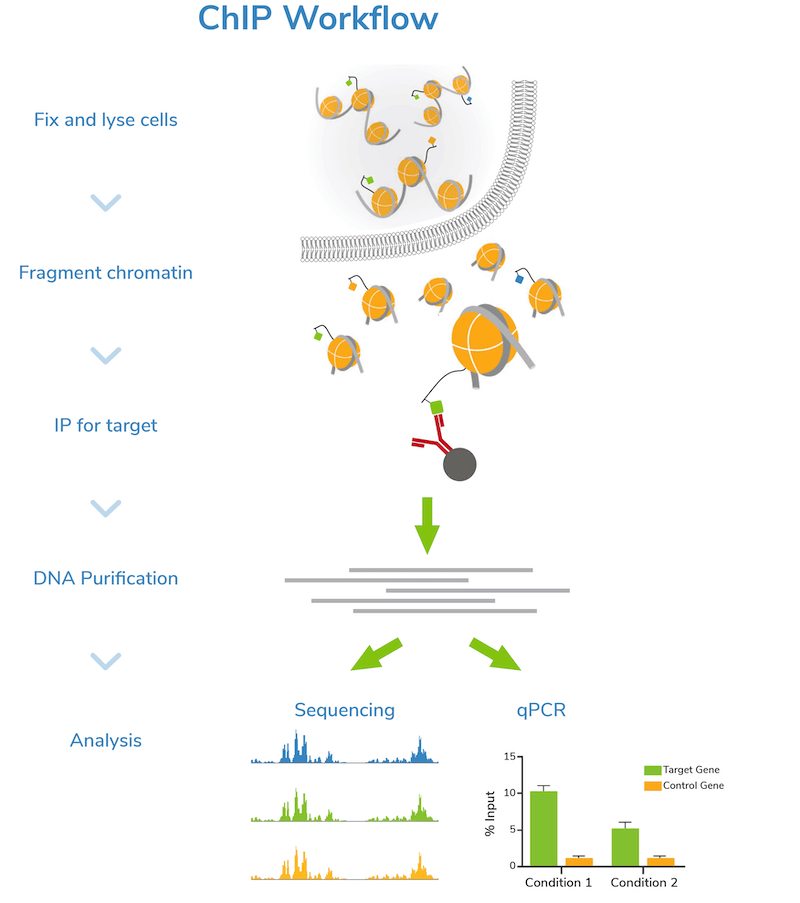
Step 2: Fragment chromatin. Next, cells are lysed, and bulk chromatin is isolated. The chromatin is fragmented or “sheared” to mononucleosome sized fragments (150-300 bp), which is important for obtaining high resolution sequencing data. Chromatin shearing is accomplished by sonication or enzymatic digestion and monitored by gel (e.g. agarose) or capillary electrophoresis (e.g. Agilent Bioanalyzer® or TapeStation®). An aliquot of fragmented chromatin is set aside as an “Input” control for quality control assessment and enrichment comparisons in later steps.
CRITICAL STEP: Chromatin shearing is the most challenging step of the ChIP workflow, requiring a delicate balance to achieve the desired level of fragmentation. See our notes on optimizing ChIP assays in the next section.
Step 3: Incubation with target-specific antibody. Sheared chromatin is incubated with a primary antibody, which binds to chromatin fragments carrying the target of interest. It is strongly recommended to include negative control reactions using an IgG antibody to assess background signal. If enough cells are available, using a validated positive control antibody (e.g. H3K4me3) is also suggested. Antibody incubation is typically performed overnight at 4˚C.
CRITICAL STEP: It is essential to use a high-quality antibody that efficiently captures its target with minimal cross-reactivity to other histone PTMs or chromatin-associated proteins. Read more about antibody selection below!
Step 4: Immunoprecipitation. The antibody is coupled to magnetic beads coated with protein-A and/or G (depending on the antibody isotype) to facilitate immunoprecipitation (Figure 1). Depending on the protocol, beads may be coupled to primary antibodies before the overnight antibody incubation, or on the following day. Regardless, the antibody-bound chromatin is isolated from bulk chromatin using a magnet, followed by a series of stringent washes. Wash buffers with progressively higher salt and detergent concentrations are used to strip away off-target proteins to reduce background signal in the data.
Step 5: DNA purification and quality check. The target-enriched chromatin is treated with Proteinase K to digest proteins, RNase A to degrade RNA, and high salt + heat to reverse cross-links. ChIP DNA is then purified using a DNA purification kit or equivalent method. DNA is also purified from the Input sample at this time.
The concentration of ChIP and Input DNA is assessed by spectrophotometric (e.g. Thermo Fisher Scientific Nanodrop™) or fluorometric (e.g. Invitrogen Qubit™) analysis. Agarose gel or capillary electrophoresis (Bioanalyzer® or TapeStation®) are used to confirm fragment size distribution prior to next steps. It is important to confirm that the ChIP DNA is enriched for mononucleosome-sized fragments, as opposed to very short or long pieces, to ensure successful downstream analysis.
Step 6: Next-generation sequencing (NGS) or qPCR. Now it’s time to analyze your sample! Sequencing allows genome-wide target studies, whereas qPCR provides a narrow look at specific region(s) of interest. The Input control is included in both strategies to determine enrichment. The IgG negative control is also analyzed to assess levels of nonspecific background signal.
For ChIP-seq studies, additional steps are required prior to sequencing. In brief:
Step 7: Library prep and indexing. ChIP DNA and Input DNA are repaired and amplified for NGS. During PCR, distinct indexes (or barcodes) are added to each unique library to allow multiplexed NGS. Multiplexing allows many samples to be sequenced in a single run.
Step 8: Library quantification and analysis. Prepared libraries are quantified and size distribution is confirmed, typically by capillary electrophoresis (Bioanalyzer® or TapeStation®).
Step 9: Sample pooling and loading onto the sequencer. For multiplexed sequencing, barcoded libraires are pooled at equimolar ratios and loaded onto the NGS platform. Time for NGS varies by platform, number of bases being sequenced, and whether paired-end sequencing is necessary.
The above steps are a simplified outline of a ChIP experiment. As any experienced chromatin scientist will tell you, ChIP is a challenging and time-consuming technique – there are multiple places for the assay to go awry.
What makes ChIP complicated? All the Optimization!
Every step of ChIP requires optimization. Furthermore, any modification to one step will likely affect downstream steps. For example, the number of cells required depends in part on your cross-linking conditions, the degree of chromatin fragmentation, as well as the specificity and enrichment efficiency of your antibody. Likewise, antibody selection impacts sequencing depth needed to achieve high signal over background. The interdependence of each step complicates assay development with iterative optimization quickly consuming significant time, money, and resources.
Below we highlight some common parameters to consider during ChIP optimization.
Number of Cells. The total number of cells needed depends on the intrinsic properties of your target, such as overall abundance, and efficacy of downstream processing steps. In general, ChIP reactions require a minimum of 500,000 cells; but in practice, most researchers use million(s) of cells per ChIP.
Replicates. Replicates are also important for ChIP, since the data can vary across reactions. You should plan for three biological replicates per target. A biological replicate represents independently collected cell or tissue samples. For instance, if you want to map H3K4me3 in intestinal epithelial cells from control mice and mutant mice models, you will want to harvest tissue from three controls and three mutant mice. Conversely, a technical replicate represents a repeated procedural step performed on the same biological sample (e.g. ChIP replicates performed using Input DNA from the same mouse)8. Technical replicates may be considered in early optimization stages to ensure robust conditions for future experiments.
Cross-linking conditions. This step requires careful optimization for your cell type and the target of interest. Parameters to adjust include the number of cells harvested as well as the cross-linking agent (e.g. formaldehyde, glutaraldehyde, disuccinimidyl glutarate [DSG]), agent concentration, and incubation time. Excessive cross-linking can mask the antibody epitope, prevent effective chromatin shearing, and degrade the sample. Alternatively, insufficient cross-linking can increase sensitivity to mechanical shearing processes, reduce total captured target chromatin, and impact data analysis. Typically, a time-course experiment is performed to determine the optimal cross-linking time and concentration.
Note on Native ChIP: For highly stable histone-DNA interactions, or for targets tightly associated with DNA (e.g. histone PTMs), cross-linking is not necessary and ChIP can be performed under native, unfixed conditions. However, in native ChIP workflows, the nuclei are first isolated from cells before fragmentation. See EpiCypher’s recently published protocol chapter for detailed native ChIP methods1.
Chromatin fragmentation. Chromatin shearing is routinely highlighted as one of the most difficult parts of ChIP experiments. Generally, a size range from 150-300 bp is desirable (i.e. between mononucleosome and dinucleosome fragments). If the pieces of chromatin are too long (>600-700 bp), it becomes difficult to identify exactly where the protein or histone PTM is located, thus lowering resolution. Furthermore, larger fragments immunoprecipitate more efficiently due to antibody avidity bias9. Large DNA fragments are also unsuitable for most next generation sequencing (NGS) platforms, which prefer genomic DNA fragment sizes of 200-600 bp. On the other hand, excessive fragmentation disrupts target interactions and reduces ChIP yields.
Fragmentation methods: In standard cross-linked ChIP workflows, bulk chromatin can be fragmented by sonication or enzymatic digestion using micrococcal nuclease (MNase). Native ChIP samples require MNase digestion, as mechanical shearing processes can disrupt unfixed chromatin and alter genomic profiles.
Optimization: For both sonication and digestion, a time course is performed to determine the ideal number of sonication cycles/digestion time, ideal ratio of buffer volume vs. number of cells, and compatibility with cross-linking conditions. It is recommended to optimize fragmentation conditions for every new input type (e.g. new cell/tissue type, varied treatment/processing steps), since cross-linking conditions and cell viscosity impact fragmentation. Even for small protocol changes, it is a good idea to check that fragmentation conditions are working as expected.
Antibody selection. The success of a ChIP experiment is dependent on using a highly specific and efficient antibody. This means that the antibody shows low binding to off-target proteins or histone PTMs, and that the antibody recovers a high percentage of the target. How should antibodies be selected?
Histone PTM antibodies notoriously show high levels of cross-reactivity in ChIP which can mislead biological conclusions1,9–13. To address this issue, EpiCypher developed SNAP-ChIP® Spike-in technology, which uses DNA-barcoded designer nucleosomes to assess histone PTM antibody performance directly in a ChIP experiment. This approach is more reliable than surrogate assays (e.g. histone peptide arrays, immunoblot, ELISA) that do not adequately mimic ChIP10. We also offer a collection of SNAP-ChIP Certified Antibodies to histone PTM targets, which have been extensively validated for high specificity and efficiency in ChIP assays. Be sure to visit chromatinantibodies.com and look up your histone PTM for detailed information on commercial antibody efficiency and specificity.
If trying to identify antibodies for chromatin-associated proteins, or if no ChIP-grade antibodies are available, it is recommended to source 3-5 antibodies from various reputable vendors and that bind distinct epitopes. Do a side-by-side comparison and select the best antibody based on ChIP yields, enrichment, and generation of expected peak structures consistent with the target function(s).
Note on lot-specific testing: Antibody performance can vary drastically lot-to-lot, even for some monoclonal antibodies. EpiCypher recommends re-testing every new lot of antibody to confirm specificity and efficiency is maintained in ChIP. If you find an excellent antibody to a target of interest, it is recommended to stock up on aliquots from the same production lot!