Highly specific antibodies are a crucial component of epigenetics research, and form the foundation of multiple chromatin profiling assays. For instance, in chromatin immunoprecipitation experiments (i.e. ChIP-seq or ChIP-qPCR), antibodies are used to map the localization of histone post-translational modifications (PTMs) and other chromatin associated proteins. The specificity of these antibodies is thus paramount to the biological relevance of resulting ChIP datasets.
But how is antibody specificity determined? Are there methods for reliable and accurate ChIP antibody validation?
ChIP Antibody Validation Strategies
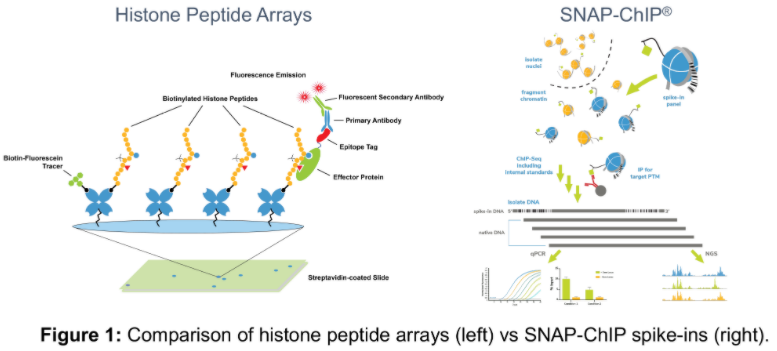
Histone peptide arrays have long been the gold standard method for selecting and validating antibodies to unique histone PTMs. These arrays test the binding capability of an antibody against a large and diverse panel of modified histone peptides, immobilized to a solid surface and subjected to wash conditions similar to Western Blot (Figure 1)1.
This strategy is fast, affordable, and contains a diverse panel of single and combinatorial PTMs.
But do histone peptide arrays accurately predict antibody binding against a physiological chromatin substrate, as in ChIP?
EpiCypher was also interested in this question, and created the SNAP-ChIP® (Sample Normalization & Antibody Profiling for ChIP) technique. This technology uses a pool of DNA-barcoded modified recombinant nucleosomes as an internal spike-in controls for ChIP (Figure 1), providing an alternative antibody validation method to histone peptide arrays2, 3.
To put SNAP-ChIP to the test, we sourced a library of >50 antibodies to H3K4 methyl states (me0, me1, me2, m3), and screened their binding specificity in histone peptide arrays and against SNAP-ChIP (using a focused set of DNA-barcoded recombinant nucleosomes)3. Based on the widespread use of modified histone peptides in the chromatin field, we expected that the majority of antibodies would behave similarly on dNucs and histone peptide substrates. But this is not what we found…
Head-head Comparison of Histone Peptides and Nucleosomes
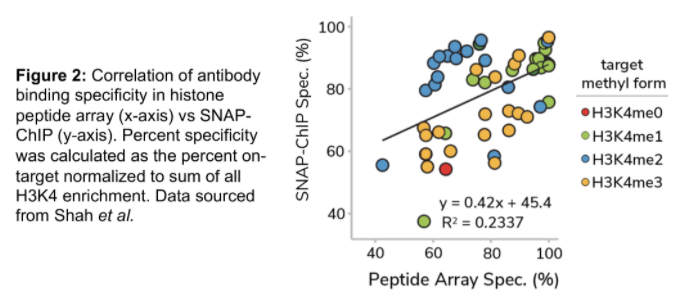
Remarkably, there was no correlation between antibody performance in SNAP-ChIP and histone peptide arrays (Figure 2). In fact, binding to histone peptides failed to predict performance of antibodies in ChIP-seq experiments!
Why Don’t Histone Peptide Arrays Work?
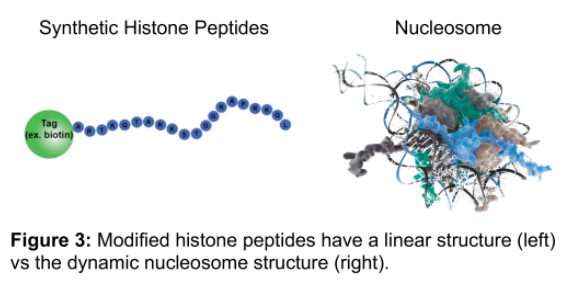
This shocking result can be most easily explained through analysis of histone peptide and nucleosome substrates. The linear epitopes represented in modified histone peptides are substantially different from the complex nucleosome structure an antibody actually interacts with in ChIP applications (Figure 3).
Widespread Problems with Histone PTM Antibody Specificity in ChIP-seq
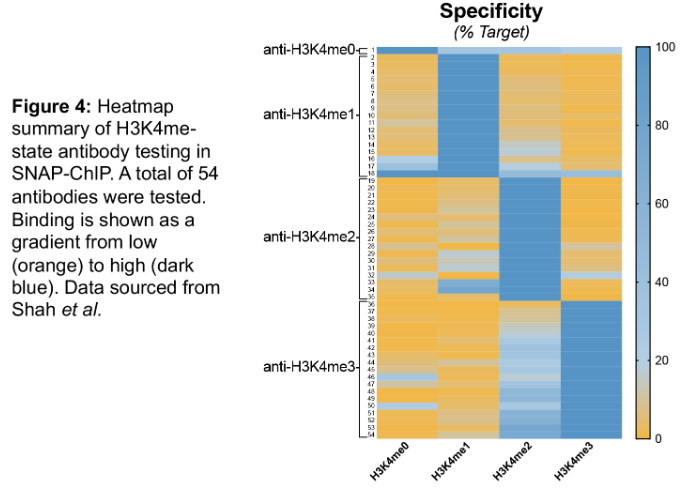
Another important conclusion from our study is that many commonly used “ChIP-grade” antibodies are not specific for their intended histone PTM target (Figure 4). However, SNAP-ChIP was able to identify a highly specific and efficient (i.e. enrichment from Input) antibody for each PTM tested, indicating that:
1. Good antibodies to histone PTMs ARE available
2. In-application testing against a defined nucleosome substrate, as in SNAP-ChIP, is the ideal strategy to identify highly specific antibodies
The Consequences of Poor Antibody Validation
While analyzing our SNAP-ChIP dataset, we were particularly dismayed by the results for H3K4me3 antibodies : of the 19 H3K4me3 antibodies tested, 16 exhibited >10% cross-reactivity with H3K4me2 (see Figure 5).
This is particularly alarming, as H3K4me2 is 3-5 times more abundant in the cell compared to H3K4me3. In addition, this low-specificity antibody group included many of the most highly cited H3K4me3 antibodies, and have been used in widely cited ChIP-seq datasets.
Thus, much of the binding that has been attributed to H3K4me3 in previous studies may actually be contaminating H3K4me2 signal!
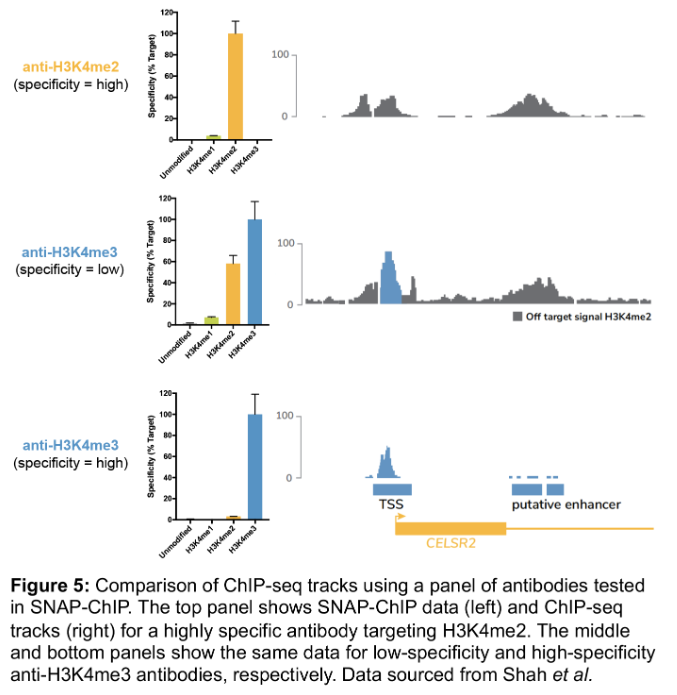
We wanted to test this theory, so we performed ChIP-seq experiments using three different antibodies (Figure 5):
1. A H3K4me2 antibody with high specificity
2. A H3K4me3 antibody with binding to both me3 and me2 marks, that has >500 citations!
3. A H3K4me3 antibody that only binds me3
Comparison of ChIP tracks (Figure 5) revealed that use of a low specificity H3K4me3 antibody results in significant contaminating signal from H3K4me2.
This finding has important biological implications. In addition to the classical association of H3K4me3 with active promoters, H3K4me3 has also been described at actively transcribed enhancers, and “broad domains” of low-level H3K4me3 enrichments have been associated with expression of genes dictating cell identity.
However, after correcting for background H3K4me2 and me1 signal using SNAP-ChIP, none of these results were replicated. It is likely that many of the biological functions assigned to H3K4me3 may be due to the use of poorly validated antibodies.
Thus, it is essential to properly validate histone PTM antibodies against a nucleosome substrate in the context of a ChIP experiment. Of note, SNAP-ChIP is the first and only platform that allow you to examine antibody specificity during your ChIP experiment.
Moving Forward : How to Choose an Antibody for ChIP?
The use of defined nucleosome substrates in SNAP-ChIP is essential for accurate testing of histone PTM antibody activity. SNAP-ChIP spike-ins are also an invaluable tool for monitoring experimental variation, and provide a STOP / GO checkpoint prior to sequencing.
EpiCypher is committed to improving the reliability and accuracy of histone PTM antibodies for chromatin research. Our SNAP-ChIP Certified Antibodies, spike-in panels, and antibody validation services identify high performance ChIP-grade antibodies to generate quality ChIP-seq data you can trust.
In addition, we have recently partnered with ThermoFisher Scientific to deliver best-in-class SNAP-ChIP Certified Antibodies. This collection includes antibodies targeting histone methylation and acetylation, and is being constantly updated as a result of our continued R&D efforts.
Stay tuned for advice on how to choose an antibody for your next ChIP experiment! For more information, send us a note at info@stratech.co.uk, and follow us on Twitter for the latest in chromatin research.
REFERENCES
1. Rothbart SB, et al. Peptide microarrays to interrogate the “histone code”. Methods Enzymol, 2012. 512: p. 107-35. (PubMed PMID: 22910205) (PMC3741997)
2. Grzybowski AT, et al. Calibrating ChIP-Seq with Nucleosomal Internal Standards to Measure Histone Modification Density Genome Wide. Mol Cell, 2015. 58(5): p. 886-99. (PubMed PMID: 26004229) (PMC4458216)
3. Shah RN, et al. Examining the Roles of H3K4 Methylation States with Systematically Characterized Antibodies. Mol Cell, 2018. 72(1): p. 162-77 e7. (PubMed PMID: 30244833) (PMC6173622)