| Description | Catalogue# (links to products) | Expression system | Sequence | Tag |
| Human ACE2 | bs-46001P; bs-46002P | HEK293 | Gln18- Ser740 | His-Avi or Fc-Avi |
| SARS-CoV-2 Spike S | bs-46007P | HEK293 | Gln14-Trp1212 | Fc-Avi |
| SARS-CoV-2 Spike RBD | bs-46003P; bs-46003P-Biotin; bs-46005P | HEK293 | Arg319-Asn532 | His-Avi or Fc-Avi |
| SARS-CoV-2 Spike S1 | bs-46004P; bs-46004P-Biotin; bs-46006P | HEK293 | Gln 14-Arg 683 | His-Avi or Fc-Avi |
| SARS Spike RBD | bs-46008P; bs-46008P-Biotin | HEK293 | Arg306-Phe527 | His-Avi |
| SARS Nucleocapsid Protein | bs-49002P | E. coli | full-length | n.a. |
Bioss COVID-19
Research Solutions for Coronavirus – Bioss
Products
| Target | Type | Catalogue# | Reactive Species |
| ACE2 | Polyclonal (Rabbit) | bs-23027R; bs-23028R; bs-23443R; bs-23444R | Human, Mouse, Rat, etc |
| ACE2 | Monoclonal (Rabbit) | bsm-52614R | Human, Mouse, Rat |
| Nucleocapsid Protein | Monoclonal (Mouse) | bsm-41411M; bsm-41412M; bsm-41413M; bsm-41414M; bsm-41415M | SARS-CoV-2 |
| Nucleocapsid Protein | Polyclonal (Rabbit) | bs-41408R | SARS-CoV-2 |
| Spike Glycoprotein | Polyclonal (Rabbit) | bs-0130R; Conjugated Versions | SARS |
| Spike Glycoprotein S1 | Polyclonal (Rabbit) | bs-17239R | SARS |
| ORF1a Polyprotein | Polyclonal (Rabbit) | bs-0132R; Conjugated Versions | SARS |
| Nucleocapsid Protein | Monoclonal (Mouse) | bsm-49131M; bsm-49132M; bsm-49133M; bsm-49134M; bsm-49135M | SARS |
| Nucleocapsid Protein | Polyclonal (Rabbit) | bs-49002R | SARS |
| Nucleocapsid Protein | HRP-conjugated | bsm-49131M-HRP; bsm-49133M-HRP | SARS |
Covid-19 Related Bioss Blogs
COVID-19 Vaccines being Tested in Clinical Trials
The coronavirus juggernaut continues to travel across the globe. For now, we are relying on tools such as physical distancing to control the spread of the virus, but in the long term a COVID-19 vaccine may be our best hope for a return to normal life. Therefore, the progress of a COVID-19 vaccine is perhaps being watched the closest by the general public.
Although creating a safe vaccine for a new illness is no easy feat, rapid progress is still being made for a variety of reasons, including China’s efforts to sequence the genetic material of SARS-CoV-2 and to share that information with research groups around the world. Another factor contributing to the unprecedented speed of development is the fact that other coronaviruses such as SARS and MERS, were already on the radar of health science researchers and learnings from them have been applied to defeating COVID-19. As of 12 April 2020, among COVID-19 vaccines being studied in different clinical phases, we summarized the most promising candidates and listed their essential information in Table 1.
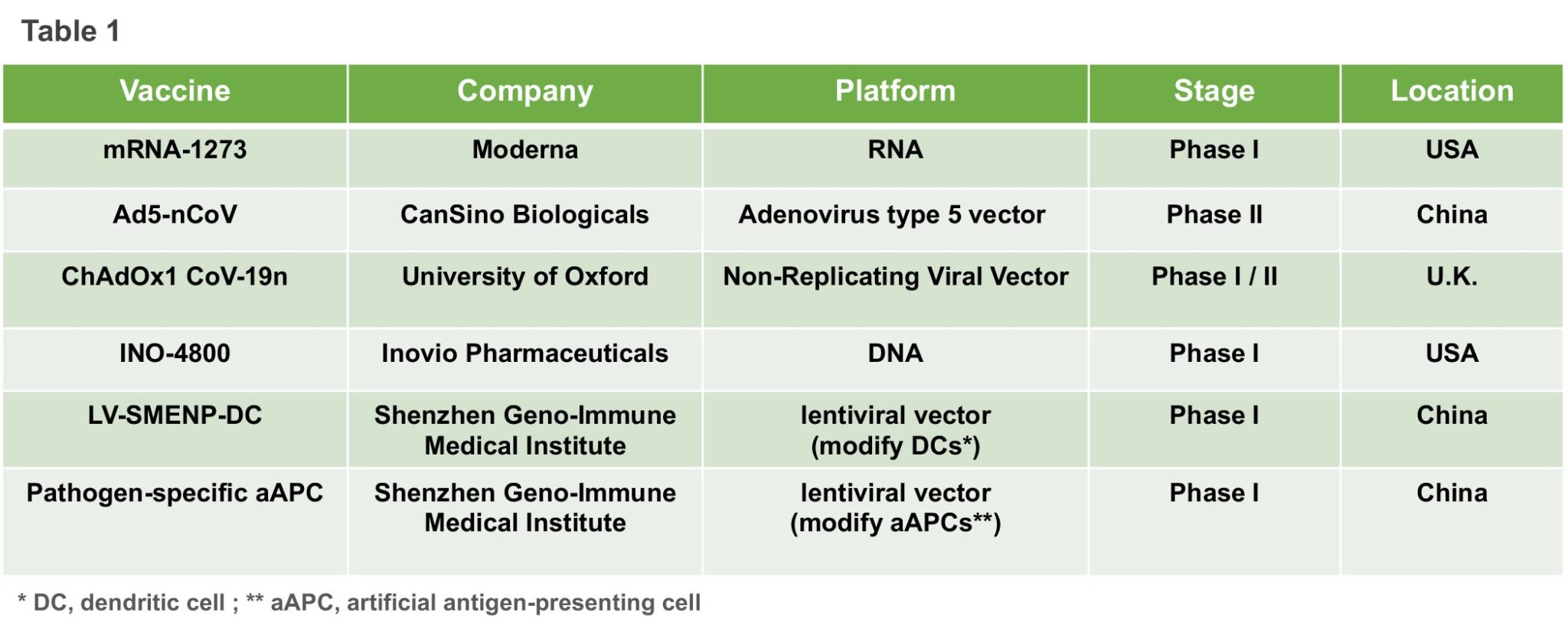
When we were preparing this summary, China has approved two more early-stage human vaccine tests for COVID-19. The experimental vaccines are being developed by a Beijing-based unit of Sinovac Biotech and by the Wuhan Institute of Biological Products, an affiliate of state-owned China National Pharmaceutical Group, state news agency Xinhua reported on Tuesday. However, more detailed technology information remains limit.
Anthony Fauci, member of the White House coronavirus task force and director of the National Institute of Allergy and Infectious Diseases, and other top U.S. health experts have previously stated that it could take researchers 18 months to create a working vaccine. Meanwhile, many health providers also concerned that, even if an effective vaccine is created shortly, it would be difficult to massively produce and apply it in a timely manner. Undoubtedly, strong international coordination and cooperation between vaccine developers, regulators, policymakers, public health bodies, and governments will be needed to ensure that promising late-stage vaccine candidates can be manufactured in sufficient quantities and equitably supplied to all affected areas, particularly low-resource regions.
A brief overview of COVID-19 diagnostic tests
COVID-19 is the infectious disease caused by a novel coronavirus, which was first identified by Chinese authorities and provisionally named as 2019-nCoV. The International Committee on Taxonomy of Viruses gave the official name SARS-COV-2 to the novel coronavirus shortly after. SARS-COV-2 has a close genetic similarity with SARS-CoV, the other viral agent identified from another coronavirus outbreak in 2003. From the previous epidemic, we learned that the rapid and widespread point-of-care (POC) testing is the foundation to curb the coronavirus pandemic.
“So far, the frontline response to the SARS-CoV-2 outbreak has been polymerase chain reaction (PCR) testing. PCR is the gold standard for diagnosing an infectious agent, and it has the advantage that the primers needed for such tests can be produced with relative speed as soon as the viral sequence is known.” An article that was first published on March 24, 2020, mentioned. Currently, large providers such as Roche Diagnostics, Thermo Fisher Scientific, Qiagen (soon to be acquired by Thermo Fisher), and Quest Diagnostics are ramping up the capacity to perform such tests by rolling out automated SARS-CoV-2 testing systems and services.
Notably, the PCR test is complicated and expensive and is mainly suited to large, centralized diagnostic laboratories. Tests typically take 4–6 hours to complete, but the logistical requirement to ship clinical samples means the turnaround time is 24 hours at best. To accelerate COVID-19 testing, immunoassays based on antibody-antigen recognition drew public health authorities’ attention. Either by using cloned viral antigens to detect patient antibodies directed against the virus or by using monoclonal antibodies (mAbs) to detect viral antigens in clinical samples, immunoassays can provide historical information about viral exposure, as well as diagnostic evidence.
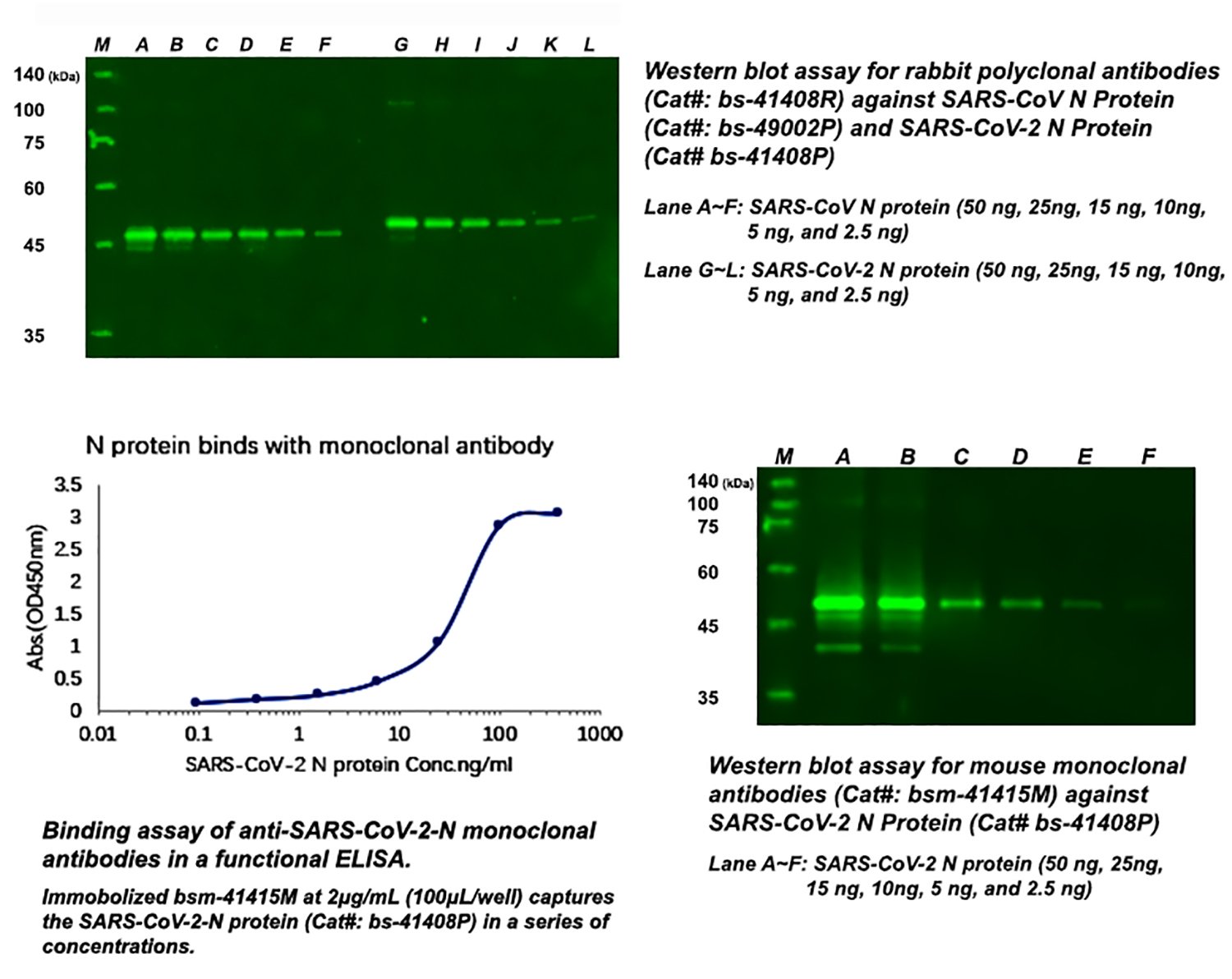
Berlin-based Pharmact has designed a 20-minute immunoassay to detect any patient antibody that recognizes three SARS-CoV-2 antigen proteins: the nucleocapsid (N) protein and the S1 and S2 domains of the spike (S) protein. This assay detects both immunoglobulin (Ig) M and IgG antibodies, which are released during the initial and later stages of an infection, respectively. It has conducted validation studies that compared its test with PCR, using samples from 114 infected patients and 126 uninfected controls. The test obtained zero false positives, indicating the perfect assay specificity. However, its sensitivity was lower, as the IgM response does not offer a strong initial signal. During the early stage of the infection (days 4–10), the IgM component of the test provides a sensitivity of just 70%. This rises rapidly to 92.3% between days 11 and 24, and the IgG component of the test offers a sensitivity of 98.6% during this phase of the infection. Overall, the test has a false-negative rate of 13%.
In parallel to tests to detect polyclonal antibodies against SARS-CoV-2 in patients, the other immunoassay is to detect the virus itself. A research team in Taiwan claimed to be first to develop a mAb against the N protein of SARS-CoV-2, which could form the basis of a rapid antigen test. Sona Nanotech, in cooperation with GE Healthcare Life Sciences, is also working on the assay development by targeting the S1 domain of the SARS-CoV-2 spike protein. If successfully manufactured and validated, the rapid immune-based test kit could detect coronavirus antigen in only 20 minutes without instrumentation and trained personnel. Although the immunoassay is less accurate compared to PCR test, its speed and versatility make it an invaluable test, and efforts to produce them on a massive scale are beginning to ramp up.
Self-diagnosis for COVID-19
The novel coronavirus has infected almost one million people and killed tens of thousands on nearly every continent. Even with offices shut down, people staying at home and hospitals bracing for an influx of patients, the pandemic continues its spread. Moreover, scientists from Columbia University frustratingly found that infected people with no significant symptoms helped fuel the COVID-19 pandemic. “These undocumented infections often experience mild, limited or no symptoms and hence go unrecognized, and, depending on their contagiousness and numbers, can expose a far greater portion of the population to virus than would otherwise occur,” the authors said.

Given that there is no specific medicine to prevent or treat COVID-19, most countries in the world have realized that the most effective way to contain the pandemic spread is the quarantine of all infected people. However, limited medical resources cannot fulfill all COVID-19 testing and leave no one missed, especially for people without symptoms. Right now, the majority of COVID-19 tests are real-time polymerase chain reaction tests (RT-PCR) to detect viral nucleotide. Nucleic acid tests are relatively accurate, and the latest assay platform can obtain results quickly, making the detection a real point-of-care (POC) test. However, trained medical personnel and specialized lab devices are needed. Another POC test called antibody tests was also developed to screen infected individuals. Patients with infection may generate detectable pathogen-specific antibodies (IgM and IgG) after 7-10 days. Therefore, blood samples showing positive IgG or IgM to the novel coronavirus indicate infections. Notably, as immune responses are variable in different individuals, it’s not simply to say that the negative result from antibody tests indicates “no infection”. It is conceivable that antibody tests are more suitable to check the post-infection status rather than to screen for suspected infections.
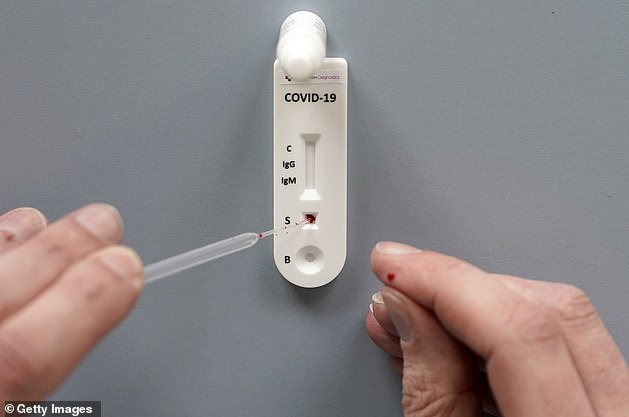
With the rapid increase of coronavirus infections, many healthcare systems are almost one step away from collapse. POC tests that do not need special instruments and trained personals will dramatically help our doctors to focus more of their time on taking care of real COVID-19 patients. Antigen tests that came out recently can directly detect viral antigens and are designed for home use. Suspected individuals can collect samples and run the test by themselves. It works kind of like a pregnancy test. The entire process takes 15 minutes, and no trained personnel or special devices are needed. This test is dependent on high-quality monoclonal antibodies to a part of the COVID-19 virus. In the face of the global pandemic, many research-driven biotech companies foresee the demand and have successfully manufactured the corresponding antibodies. Their efforts make the antigen tests available and eventually benefit our human society.
Newly Announced Epitopes of SARS-CoV-2 May Facilitate the Vaccine Design
AMERICAN SCIENTISTS IDENTIFY POTENTIAL TARGETS FOR IMMUNE RESPONSES TO NOVEL CORONAVIRUS
Within two months, SARS-CoV-2, a previously unknown coronavirus, has raced around globe, infecting over a 100,000 people with numbers continuing to rise quickly. Effective countermeasures require helpful tools to monitor viral spread and understand how the immune system responds to the virus.
Publishing in the March 16, 2020, online issue of Host, Cell and Microbe, a team of researchers at La Jolla Institute for Immunology, in collaboration with researchers at the J. Craig Venter Institute, provides the first analysis of potential targets for effective immune responses against the novel coronavirus. The researchers used existing data from known coronaviruses to predict which parts of SARS-CoV-2 are capable of activating the human immune system.
When the immune system encounters a bacterium or a virus, it zeroes in on tiny molecular features, so called epitopes, which allow cells of the immune system to distinguish between closely related foreign invaders and focus their attack. Having a complete map of viral epitopes and their immunogenicity is critical to researchers attempting to design new or improved vaccines to protect against COVID-19, the disease caused by SARS-CoV-2.
“Right now, we have limited information about which pieces of the virus elicit a solid human response,” says the study’s lead author Alessandro Sette, Dr. Biol. Sci, a professor in the Center for Infectious Disease and Vaccine Research at LJI. “Knowing the immunogenicity of certain viral regions, or in other words, which parts of the virus the immune system reacts to and how strongly, is of immediate relevance for the design of promising vaccine candidates and their evaluation.”
While scientists currently know very little about how the human immune system responds to SARS-CoV-2, the immune response to other coronaviruses has been studied and a significant amount of epitope data is available.
Four other coronaviruses are currently circulating in the human population. They cause generally mild symptoms and together they are responsible for an estimated one quarter of all seasonal colds. But every few years, a new coronavirus emerges that causes severe disease as was the case with SARS-CoV in 2003 and MERS-CoV in 2008, and now SARS-CoV-2.
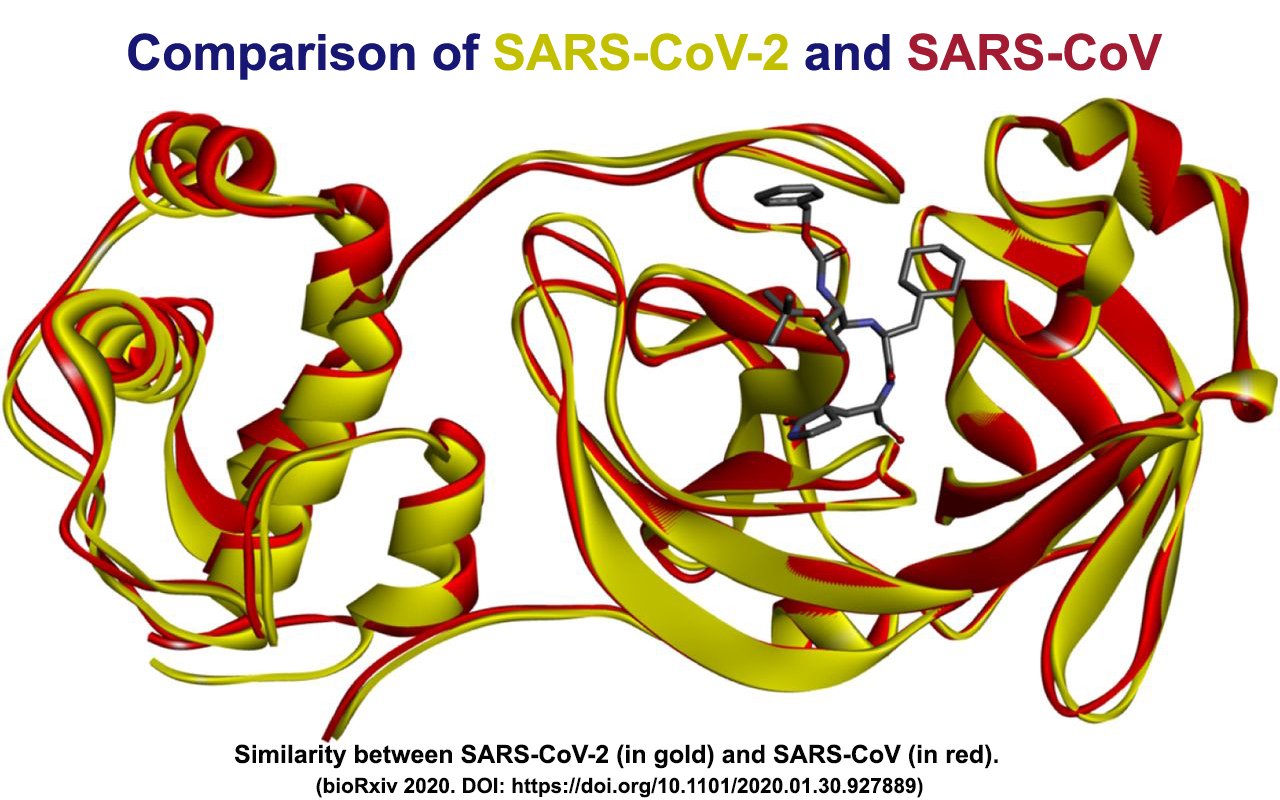
“SARS-CoV-2 is most closely related to SARS-CoV, which also happens to be the best characterized coronavirus in terms of epitopes,” explains first author Alba Grifoni, Ph.D, a postdoctoral researcher in the Sette lab.
For their study, the authors used available data from the LJI-based Immune Epitope Database (IEDB), which contains over 600,000 known epitopes from some 3,600 different species, and the Virus Pathogen Resource (ViPR), a complementary repository of information about pathogenic viruses. The team compiled known epitopes from SARS-CoV and mapped the corresponding regions to SARS-CoV-2.
“We were able to map back 10 B cell epitopes to the new coronavirus and because of the overall high sequence similarity between SARS-CoV and SARS-CoV-2, there is a high likelihood that the same regions that are immunodominant in SARS-CoV are also dominant in SARS-CoV-2 is,” says Grifoni.
Five of these regions were found in the spike glycoprotein, which forms the “crown” on the surface of the virus that gave coronaviruses their name; two in the membrane protein, which is embedded in the membrane that envelopes the protective protein shell around the viral genome and three in the nucleoprotein, which forms the shell.
In a similar analysis, T cell epitopes were also mostly associated with the spike glycoprotein and nucleoprotein.
In a completely different approach, Grifoni used the epitope prediction algorithm hosted by the IEDB to predict linear B cell epitopes. A recent study by scientists at the University of Texas Austin determined the three-dimensional structure of the spike proteins, which allowed the LJI team to take the protein’s spatial architecture into account when predicting epitopes. This approach confirmed two of the likely epitope regions they had predicted earlier.
To substantiate the SARS-CoV-2 T cell epitopes identified based on their homology to SARS-CoV, Grifoni compared them with epitopes pinpointed by the Tepitool resource in the IEDB. Using this approach, she was able verify 12 out of 17 SARS-CoV-2 T cell epitopes identified based on sequence similarities to SARS-CoV.
“The fact that we found that many B and T cell epitopes are highly conserved between SARS-CoV and SARS-CoV-2 provides a great starting point for vaccine development,” says Sette. “Vaccine strategies that specifically target these regions could generate immunity that’s not only cross-protective but also relatively resistant to ongoing virus evolution.”
———————————————————————————————–
Full citation:
Alba Grifoni, John Sidney, Yun Zhang, Richard H Scheuermann, Bjoern Peters and Alessandro Sette. A Sequence Homology and Bioinformatic Approach Can Predict Candidate Targets for Immune Responses to SARS-CoV-2. Cell, Host and Microbe, 2020.
https://doi.org/10.1016/j.chom.2020.03.002
The work was funded in part by the National Institute of Allergy and Infectious Diseases, a component of the National Institutes of Health through contracts 75N9301900065, 75N93
————————————————————————————————-
Story Source:
Materials provided by La Jolla Institute for Immunology. Note: Content may be edited for style and length.
